 
"Biochemistry"(1st ed.) Harper&Row,Publishers, Inc.pp. T7 DNA replisome by Marius Mihasan References It binds to RNA polymerase, RNA and DNA at the pause signal and RNA is released.įor additional information, see: DNA Replication, Repair, and Recombinationįor additional information, see: Nucleic Acidsįor additional information, see: Translation Files for 3D printer The termination process is aided by a protein called rho factor. When RNA polymerase reaches these T rich sequences it pauses and the synthesized RNA is released. Specific DNA sequences rich in T followed by G-C rich sequences are responsible for termination of transcription. The binding is followed by the separation of DNA strands and synthesis of RNA by RNA polymerase. In general these regions contain the following sequence These sites are known as promoter regions. Binding at certain sites is stronger than the rest and causes the initiation of RNA synthesis. There are many binding sites on DNA for RNA polymerase. In eukaryotes there are three types of RNA polymerases called RNA polymerase 1,2 and 3 which synthesize different types of RNA. This enyme is responsible for binding RNA polymerase holoenzyme to specific transcritption sites on DNA. A sixth subunit called sigma subunit binds to core enzyme to give the holoenzyme. The RNA polymerase of prokaryotes is an oligomer composed of fice subunits arranged in an oligomer known as core enzyme. RNA polymerase binding to specific site on double-helical DNA The transcription step that involves the synthesis of mRNA from can explained in the following stages. 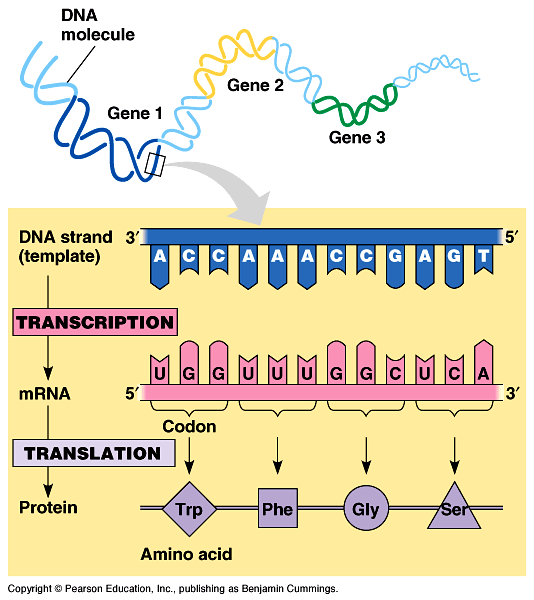
This messenger RNA is then translated into proteins on ribosomes. In the transcription stage a strand of DNA molecule serves as a template for the synthesis of an RNA molecule called messenger RNA. The expression of genes into proteins and is a process involving two stages called transcription and translation. In this step a ribonuclease removes the RNA primer, the DNA polymerase fills the gap and DNA ligase fills the nicks between the DNA fragments. Primer Excision and Phosphodiester bond formation Short strands of polynucleotides known as Okazaki fragments are formed complementary to the lagging strand. On the other strand known as lagging strand chain growth occurs discontinuously. However the synthesis occurs from 5'-3' continuously on only one strand called the leading strand. 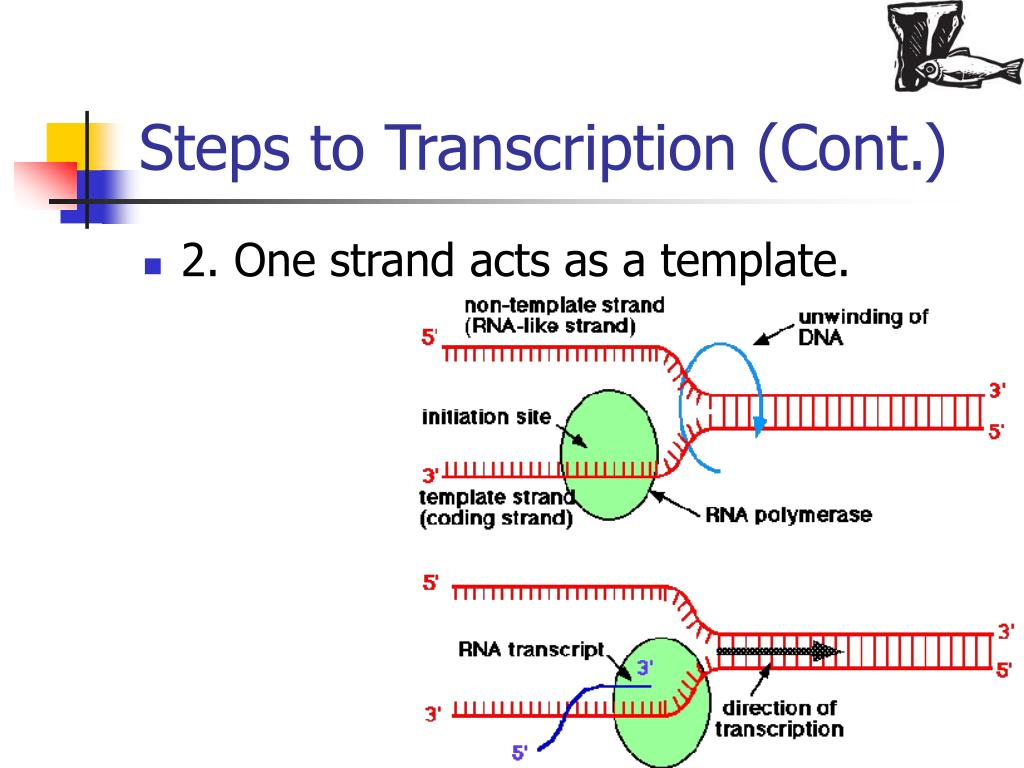
Synthesis of complementary DNA occurs simultaneously on both the parent strands. The elongation of the primer on the template strands is done by an enzyme known as DNA polymerase. This binding acts as a signal for a protein called primase an RNA polymerase which synthesizes a short complementary strand at the origin of replication.ĭiscontinuous synthesis at the Replication fork The unwinding of DNA is followed by the binding of protein dnaB at the origin of replication of DNA and remains bound to the replication fork. Another helicase called the rep protein also helps in the denaturation of DNA by disrupting the base pairs. This followed by the negative supercoiling of DNA by DNA gyrase which lowers the the energy required to disrupt a base pair. 
DNA helicase promotes the unwinding by binding to the single-stranded DNA. The supercoiled duplex DNA unwinds with the aid of several proteins. The replication can be explained in the following stages. DNA undergoes what is known as semi conservative mode of replication wherein the daughter DNA contains one DNA strand of the parent. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
